TASTE
TASTE combines the PARAFAC2 model with non-negative matrix factorization to model a temporal and a static tensor. It performs two import tasks in healthcare: 1- computational phenotyping 2- Predictive modeling by analyzing electronic health records (EHRs).
Yanxin Ye processed the CMS data using Hive.
Xinze Wang did case-control mapping using Pymatch.
Xingchi Li translated MATLAB code of TASTE and Nonnegative matrix factorization (NMF) algorithms based on alternating non-negativity constrained least squares to Python then did compatibility modification.
Hung-Yi Li did the cross validation and numerous experiments to do better comparison.
Previous TASTE code can be found in legacy which is implemented in MATLAB by Ardavan (Ari) Afshar.
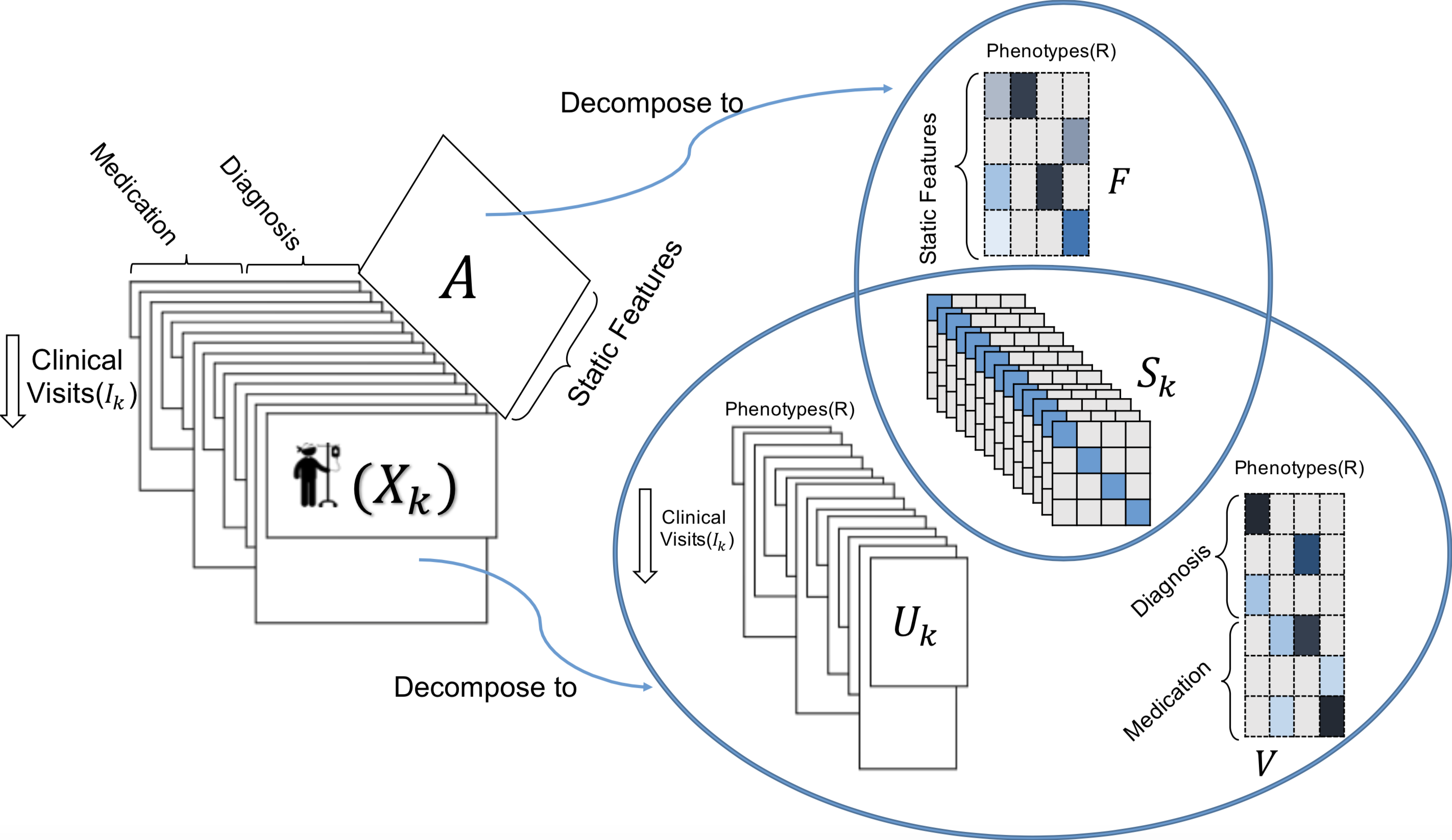
TASTE applied on dynamically-evolving structured EHR data and static patient information. Each represents the medical features recorded for different clinical visits for patient
. Matrix
includes the static information (e.g., race, gender) of patients. TASTE decomposes
into three parts:
,
, and
. Static matrix
is decomposed into two parts:
and
. Note that
(personalized phenotype scores) is shared between static and dynamically-evolving features.
Relevant Publication
TASTE implements the code in the following paper:
Afshar, Ardavan, Ioakeim Perros, Haesun Park, Christopher deFilippi, Xiaowei Yan, Walter Stewart,
Joyce Ho, and Jimeng Sun. "TASTE: Temporal and Static Tensor Factorization for Phenotyping Electronic
Health Records." ACM CHIL 2020.
Code description
Nonnegative matrix factorization (NMF) algorithms based on alternating non-negativity constrained least squares has been imported and translated in nonnegfac.py.
There are two main functions defined in main.py: fit(R, A, X) and project(R, A, X, V, F, H):
Rdenotes the number of phenotypesAdenotes the static feature matrixXdenotes the temporal feature matrixV,F,Hare the matrices obtained from thefitfunction.
In Memory of Ardavan (Ari) Afshar
Ari is a good mentor in directing us with this project. It is really sad to see a good person not around any more. RIP Ari, you will be missed. Kudoboard